- Home
- :
- All Communities
- :
- Products
- :
- ArcGIS Pro
- :
- ArcGIS Pro Questions
- :
- Edit Raster Extent with NULL values
- Subscribe to RSS Feed
- Mark Topic as New
- Mark Topic as Read
- Float this Topic for Current User
- Bookmark
- Subscribe
- Mute
- Printer Friendly Page
Edit Raster Extent with NULL values
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Permalink
Last week I posted a question in This Post but it remains unanswered. If anyone is up for the challenge, I would love to explore some creative solutions to my problem.
Some details are provided in my previous post, but basically, I would like to add a single row of NULL cells to a raster.
I have explored options such as creating a larger raster and merging the two, but am worried about this because of potential imprecision I do NOT want any resampling. Please let me know if there are other methods to try! Thank you!
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Permalink
How Aggregate works—ArcGIS Pro | Documentation
supports the Extent parameter,
however, numpy can come into play using arcpy's RasterToNumPyArray to get an array from a raster, and NumPyArrayToRaster to get it back into a raster for visualization in Pro.
arr = np.arange(9, dtype='float').reshape(3,3)
arr
Out[6]:
array([[0., 1., 2.],
[3., 4., 5.],
[6., 7., 8.]])
np.pad(arr, pad_width=1, mode='constant', constant_values=np.nan)
Out[7]:
array([[nan, nan, nan, nan, nan],
[nan, 0., 1., 2., nan],
[nan, 3., 4., 5., nan],
[nan, 6., 7., 8., nan],
[nan, nan, nan, nan, nan]])You can run the aggregation within Pro or use NumPy sliding windows functionality
... sort of retired...
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Permalink
Thank you very much for your response, indeed you are an MVP!
The NumPy approach seems to be along the lines of what I am looking for. Thank you. Upon trying this, I run into a couple issues / follow-up problems:
- Error in RasterToNumPyArray: "The pixel block exceeds the maximum size allowed" when working with my full 1-km resolution global rasters. It does run with 40-km rasters, so I am thinking of splitting up my 1-km rasters into manageable slices. Does this seem appropriate?
- HOWEVER, when I go to mosaic these together, there will be that new border I created inbetween the raster slices, which I do not want anywhere other than the outside edges.
- Using NumPyArrayToRaster and getting all of the slightly extended rasters to line up with the originals for further analysis. I am unsure on how to calculate the new "lower_left_corner" parameter. Can you please provide some advice on this as well?
- I am thinking of using the lower left corner and cell size pre-extension, and adding them together, but do worry about any imprecision, as I do not want any resampling to be done.
Otherwise, this approach works great for adding a NULL padding to my rasters and I think that I can incorporate it into a loop. Looking forward to hearing back from you! Thank you!
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Permalink
Now I am not sure what you want to do with the padded array, since you if you are doing some kind of aggregation or statistic for you "chunk", you can do it without padding.
For example, assign the modal value to an array
arr = np.random.randint(0, 3, size=(3, 3), dtype=int) # some raster to aggregate
vals, cnt = np.unique(arr, return_counts=True) # example for "mode"
mode = vals[np.argmax(cnt)] # find the modal value
arr2 = np.clip(arr, mode, mode) # assign the mode to all positions
arr # original array
array([[2, 0, 0],
[2, 2, 0],
[1, 1, 0]])
arr2 # modal array
array([[0, 0, 0],
[0, 0, 0],
[0, 0, 0]])
Same thing for floats
arr = np.random.random(size=(3,3))*10
arr
array([[2.86187486, 8.12931111, 1.73058081],
[8.04929842, 1.85048209, 6.6905913 ],
[0.35932908, 4.99030038, 0.19628441]])
avg_ = np.mean(arr)
arr2 = np.clip(arr, avg_, avg_)
arr2
array([[3.87311694, 3.87311694, 3.87311694],
[3.87311694, 3.87311694, 3.87311694],
[3.87311694, 3.87311694, 3.87311694]])
But if you want the padded values "nan" is useful since you can use "nan" functions.
arr = np.random.random(size=(3,3))*10
arr2 = np.pad(arr, pad_width=1, mode='constant', constant_values=np.nan)
avg_0 = np.mean(arr)
avg_1 = np.nanmean(arr2)
arr
array([[9.64715002, 5.43028836, 4.77926339],
[0.36168469, 6.57264434, 6.7112081 ],
[1.18304099, 7.56563182, 9.97630234]])
arr2
array([[ nan, nan, nan, nan, nan],
[ nan, 9.64715002, 5.43028836, 4.77926339, nan],
[ nan, 0.36168469, 6.57264434, 6.7112081 , nan],
[ nan, 1.18304099, 7.56563182, 9.97630234, nan],
[ nan, nan, nan, nan, nan]])
avg_0 == avg_1
True
# avg_0, avg_1 : (5.803023783223992, 5.803023783223992) just to show
so let me know what you want to do with the chunk/block before you aggregate and I can deal with piecing the pieces together with numpy "slicing" and "concatenation"
... sort of retired...
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Permalink
Dear Dan,
Thank you again for your response. I really appreciate your time and effort in helping me.
My overall goal is to aggregate my 1-km global raster to 40-km resolution at a variety (>100) of different configurations. Then, I will use these aggregated rasters in a regression tree analysis and average out the results spatially to achieve a 1-km resolution resulting raster. The reason that I wanted to incorporate this padding with Null values is to create a different configuration of 1-km cells within the 40-km block, as the aggregation will take place from the bottom-left corner. Does this make sense? Please let me know if you would like further clarification on this.
As for the chunks (of the 1-km raster that is too large to be converted into a NumPy Array), ideally I would like to convert them back into rasters and mosaic them in such a manner that I end up with a single global raster with one row of Null values around the edges, not inbetween the chunks. Also, I would like this to align with the previous not-extended raster. Is this possible? Thank you so much for any other advice. I would be happy to DM you and share my data if that would be helpful! Thanks!
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Permalink
1 km or km^2 global raster? is that 510ish million cells? is the unit width, height or area?
40 km or 40 km^2 (aka, 12.7 and 3.2 million cells
I am not sure what you mean by "configurations"
In any event, the 40 unit block is the only one that needs to be padded and the padding would be dependent whether you are doing a sliding-window or a block-window view (eg increment the subarray by 1 or by a block size like 10*10)
... sort of retired...
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Permalink
It is a 1-km^2 resolution global raster. There are 187 million land area cells, likely 510ish million cells total as you said.
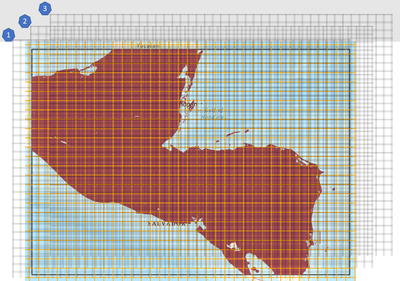
To try to explain what I mean by "configurations", I am providing the below rough graphic, where I show a spatial subset of my global data and 40-km^2 windows overlays. The configurations are the different 1-km^2 cells that are aggregated within each "shifted" 40-km^2 window. I would like to perform this shift by adding rows of padding to the 1-km^2 raster, pre-aggregation. Does this make logical sense to you?
I would like to create a loop which: 1) adds a row of padding around my 1-km^2 raster, 2) aggregates this raster to 40-km^2 resolution, 3) exports the 40-km^2 raster values to points for further analysis, and then repeats by adding another row of padding to the 1-km^2 raster. Do you know of any way that this is possible? Thank you! 🙂