- Home
- :
- All Communities
- :
- Products
- :
- ArcGIS Pro
- :
- ArcGIS Pro Questions
- :
- Weighted average using NbrWeight kernel file?
- Subscribe to RSS Feed
- Mark Topic as New
- Mark Topic as Read
- Float this Topic for Current User
- Bookmark
- Subscribe
- Mute
- Printer Friendly Page
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Permalink
Hi,
I am trying to calculate a weighted mean statistic using a neighborhood defined by a kernel file:
My kernel file defines a 3x3 cell neighborhood, with the following weights:
| 0.5 | 1 | 0.5 |
| 1 | 1 | 1 |
| 0.5 | 1 | 0.5 |
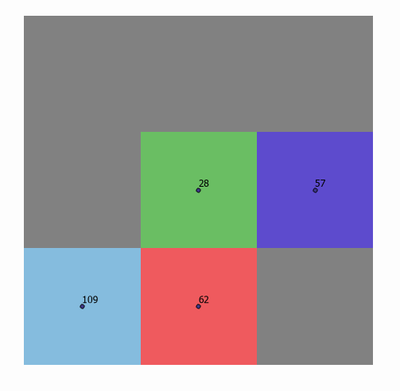
For illustration purposes, I'll use the following 3x3 raster as an example. The point labels show the values of the raster cells. Grey cells indicate no data. My target cell for this example is the center green cell, with a value of 28.
When I run the tool, the weighted mean is calculated as:
weighted sum / # of data cells in neighborhood
For the green target cell: 201.5/4 = 50.375
Output shown below:
I want it to be calculated as:
weighted sum / sum of weights of data cells in neighborhood
For the green target cell this should would be 201.5/3.5 =57.57143
How do I calculate this weighted average??
Thank you!
Solved! Go to Solution.
Accepted Solutions
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Permalink
interesting.
have you tried running the focal sum only,
produce a raster which is classed a nodata or 1 based on the presence of data in your input raster (the inverse of IsNull or a Con statement would do)
If you run a focal sum using your kernel on that raster and you should get your 3.5 value
After that, take your initial focal sum raster and divide it by your focal sum 1's raster
... sort of retired...
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Permalink
interesting.
have you tried running the focal sum only,
produce a raster which is classed a nodata or 1 based on the presence of data in your input raster (the inverse of IsNull or a Con statement would do)
If you run a focal sum using your kernel on that raster and you should get your 3.5 value
After that, take your initial focal sum raster and divide it by your focal sum 1's raster
... sort of retired...
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Permalink
Thanks Dan! This will work. I appreciate the help!
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Permalink
no problem... an interesting question
... sort of retired...