Generalization tools for rasters...
- Subscribe to RSS Feed
- Mark as New
- Mark as Read
- Bookmark
- Subscribe
- Printer Friendly Page
Scipy ndimage morphology … I am betting that is the first thought that popped into your head. A new set of tools for the raster arsenal.
But wait! Their terminology is different than that used by ArcGIS Pro or even ArcMap.
What tools are there? Lots of filtering tools, but some of the interesting ones are below
from scipy import ndimage as nd
dir(nd)[... snip ...
'affine_transform', ... , 'center_of_mass', 'convolve', 'convolve1d', 'correlate',
'correlate1d', ... 'distance_transform_edt', ... 'filters', ... 'generic_filter',
... 'geometric_transform', ... 'histogram', 'imread', 'interpolation', ... 'label',
'labeled_comprehension', ... 'measurements', ... 'morphology', ... 'rotate', 'shift',
'sobel',...]I previously covered distance_transform_edt which performs the equivalent of Euclidean Distance and Euclidean Allocation in this post
/blogs/dan_patterson/2018/10/04/euclidean-distance-allocation-and-other-stuff
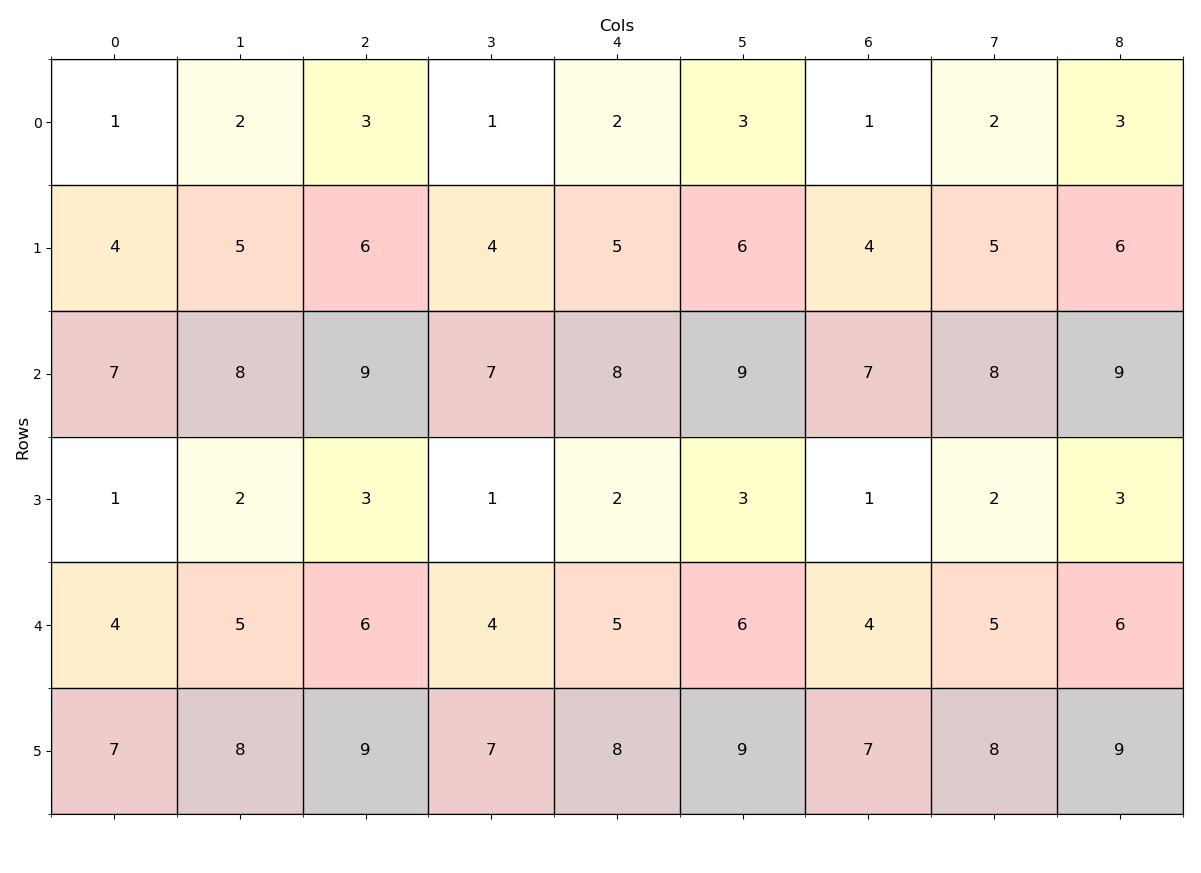
Let's begin with a raster constructed from 3x3 windows with a repeating pattern and repeats of the pattern.
You can construct your own rasters using numpy, then apply a NumPyArrayToRaster to it. I have covered this before, but here is the code to produce the raster
def _make_arr_():
"""Make an array with repeating patterns
"""
a = np.arange(1, 10).reshape(3, 3)
aa = np.repeat(np.repeat(a, 3, axis=1), 3, axis=0)
aa = np.tile(aa, 3)
aa = np.vstack((aa, aa))
return aa---- The raster (image) ----
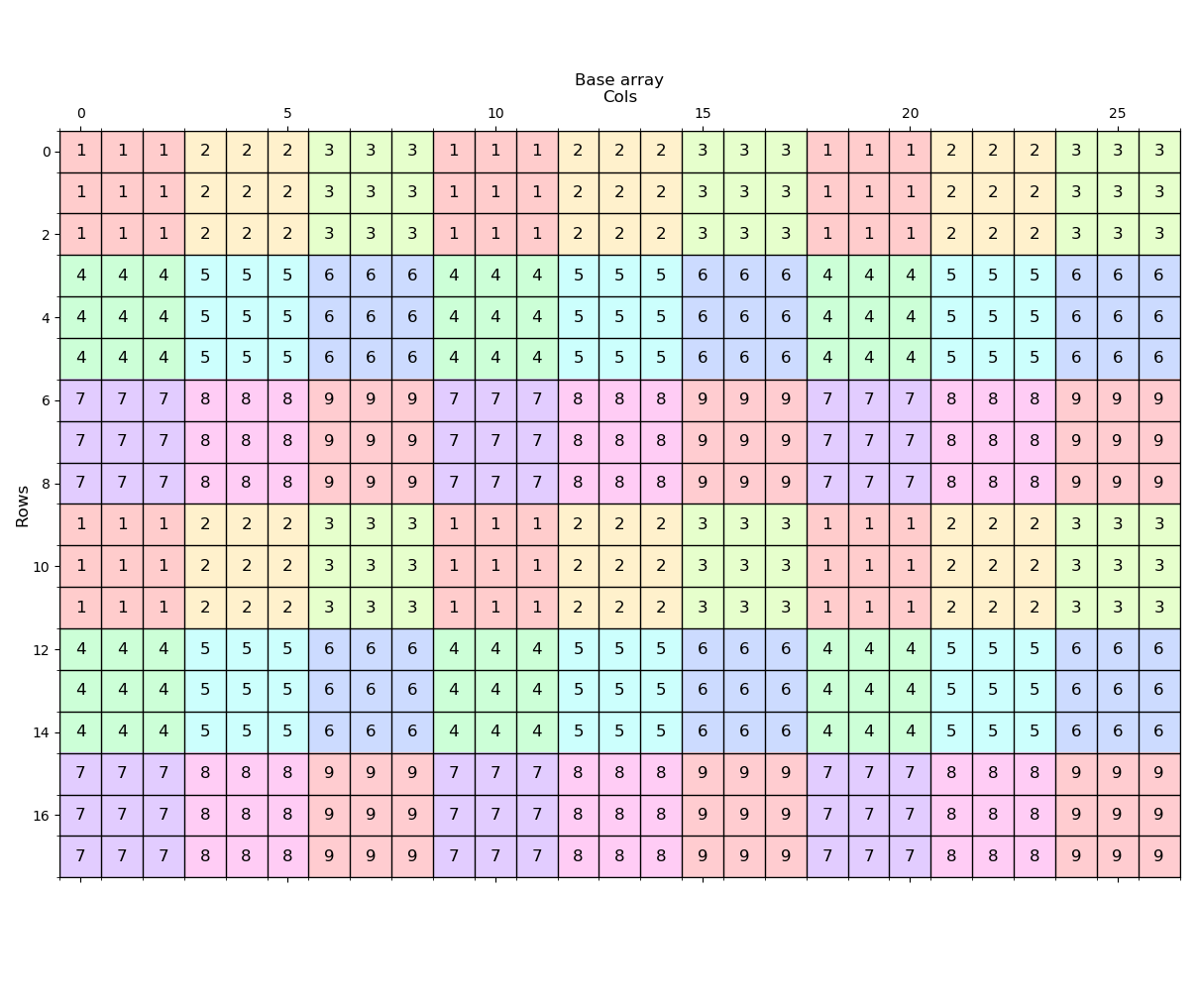
---- (1) Expand ----
Expand the zone 1 values by 2 distance units
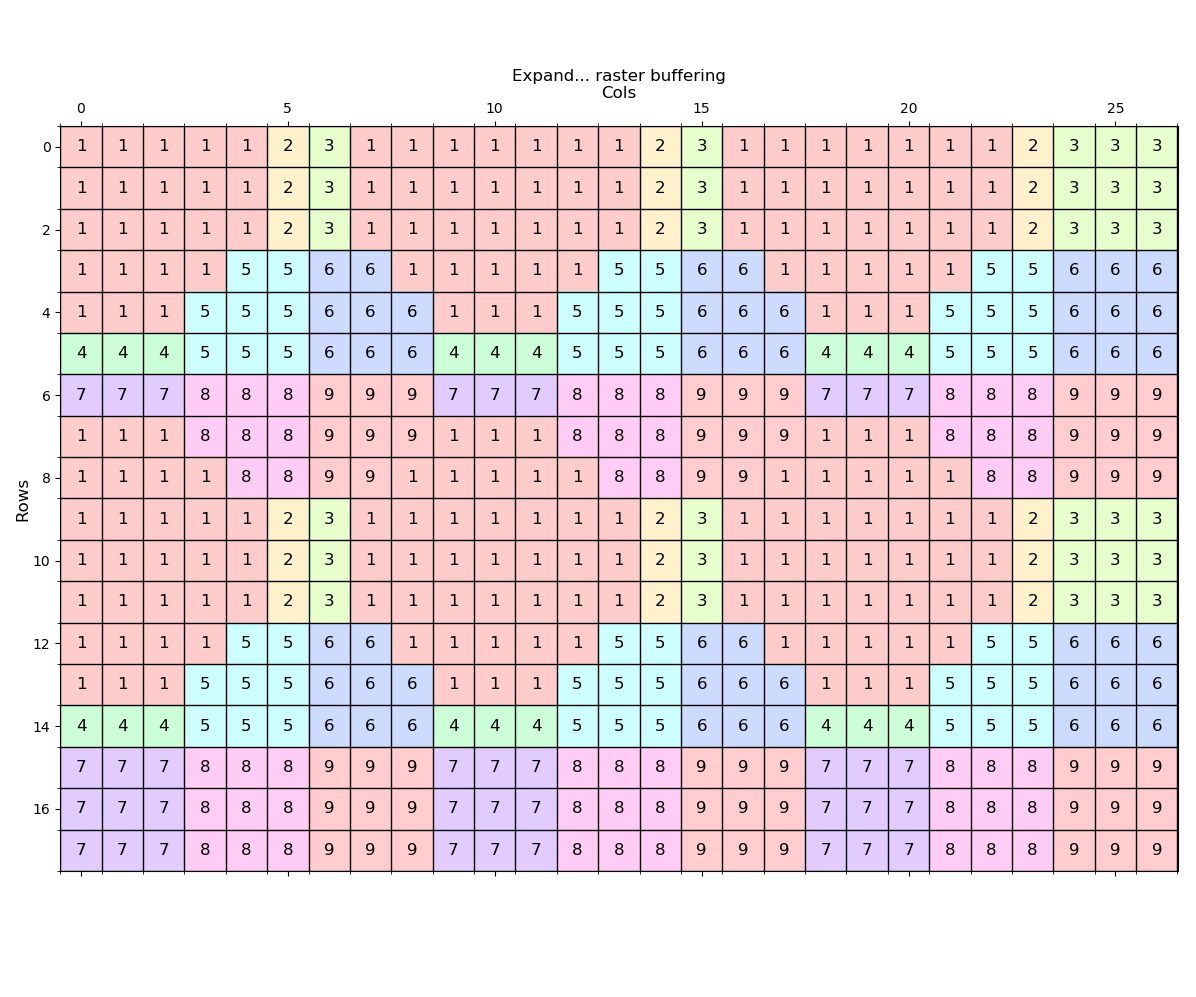
---- (2) Shrink ----
Shrink the zone 1 values by 1 distance unit.
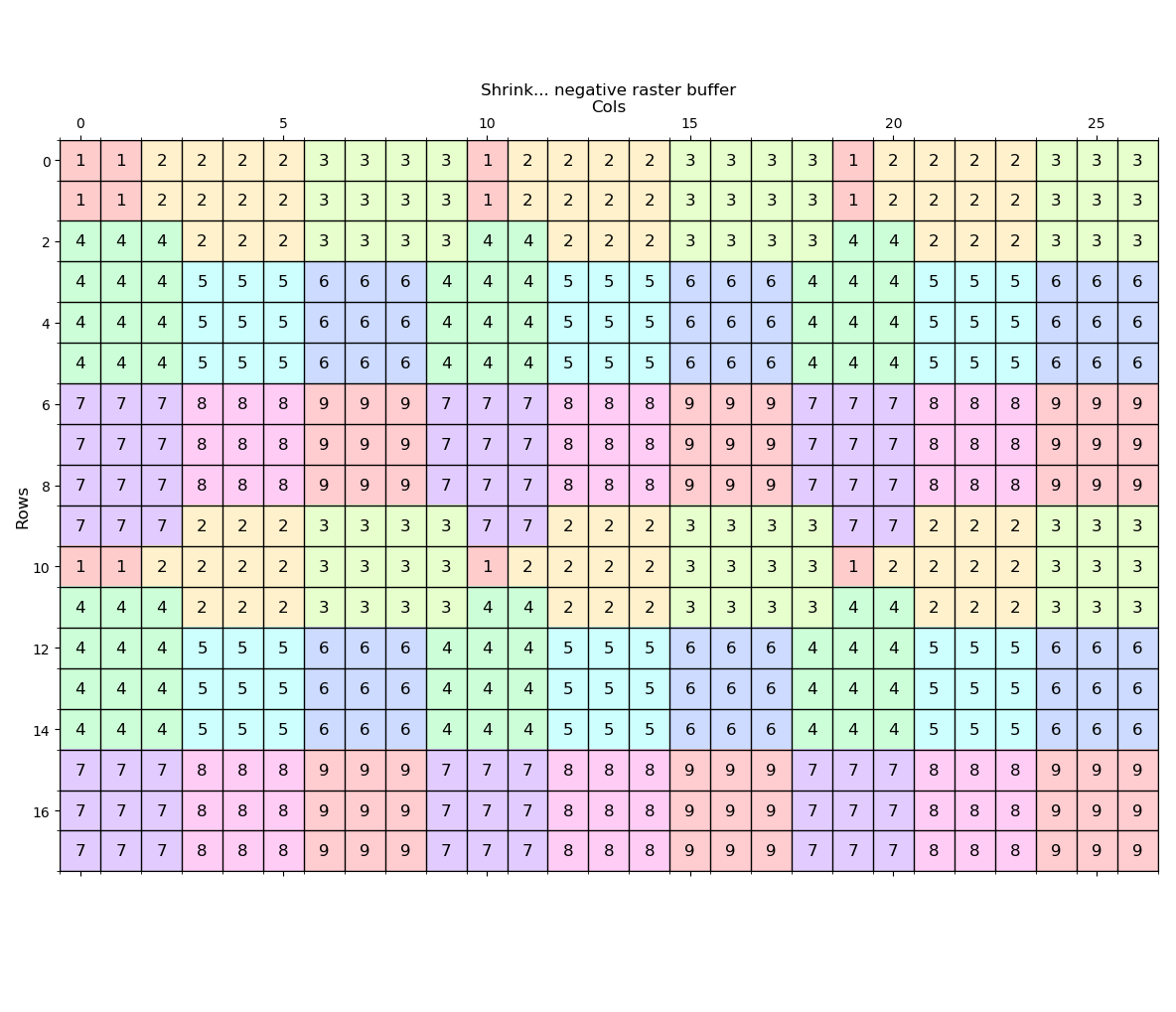
---- (3) Regiongroup ----
Produce unique regions from the zones. observe that the zone 1 values in the original are given unique values first. Since there were 6 zone 1s, they are numbered from 1 to 6. Zone 2 gets numbered from 7 to 12. and that pattern of renumbering repeats.
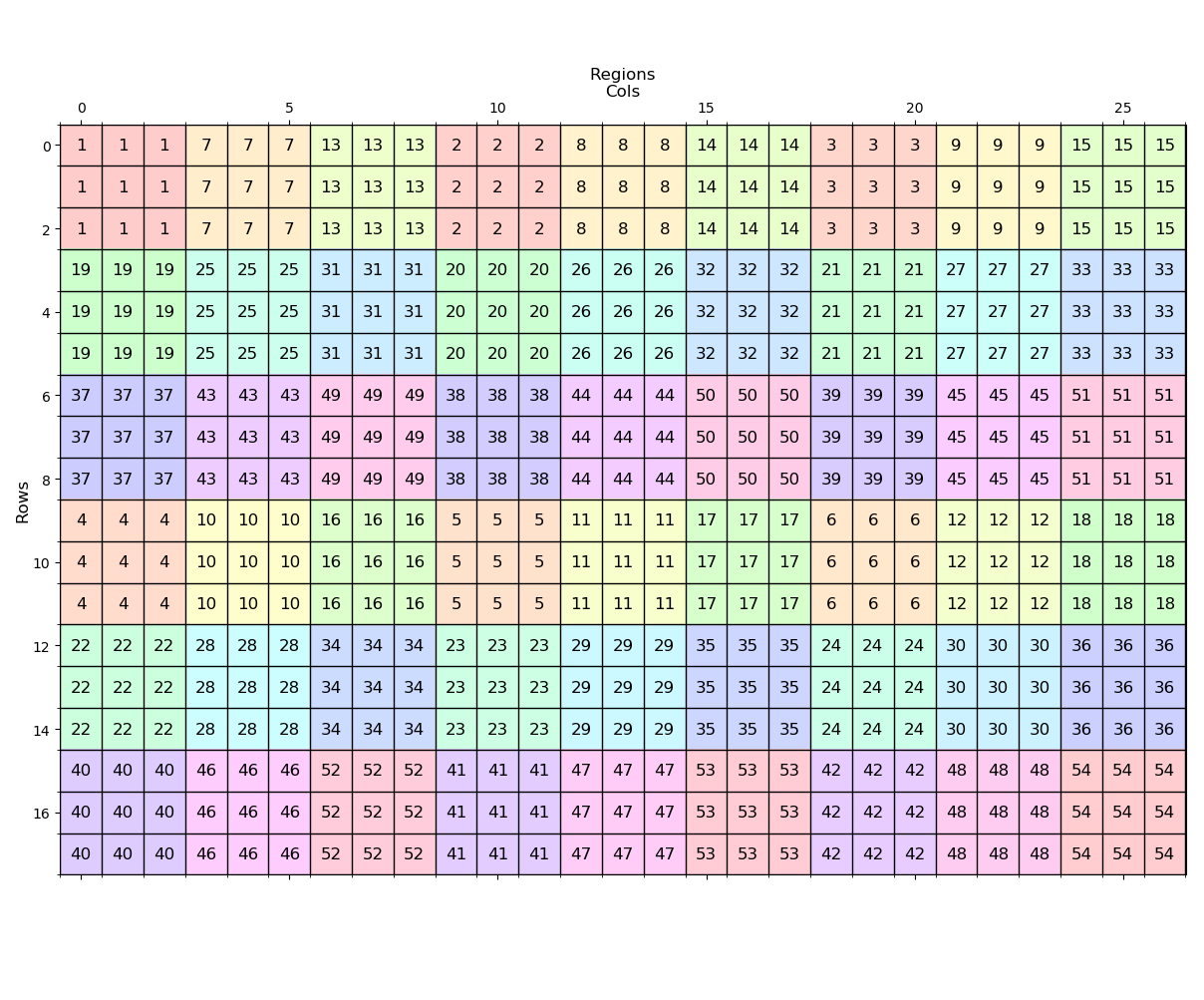
---- (4) Aggregate ----
Aggregation of the input array using the mode produces a spatial aggregation retaining the original values. Our original 3x3 kernels are now reduced to a 1x1 size which has 3 times the width and height (9x area).
---- (5) Some functions to perform the above ----
In these examples buff_dist is referring to 'cells'.
For the aggregation, there are a number of options as shown in the header.
Generally integer data should be restricted to the mode, min, max, range and central (value) since the median and mean upscale to floating point values. This of course can be accommodated by using the python statistics package median_low and median_high functions:
statistics — Mathematical statistics functions — Python 3.7.1rc1 documentation
So think of a function that you want. Filtering is a snap since you can 'stride' an array using any kernel you want using plain numpy or using the builtin functions from scipy.
Enough for now...
def expand_(a, val=1, mask_vals=0, buff_dist=1):
"""Expand/buffer a raster by a distance
"""
if isinstance(val, (list, tuple)):
m = np.isin(a, val, invert=True).astype('int')
else:
m = np.where(a==val, 0, 1)
dist, idx = nd.distance_transform_edt(m, return_distances=True,
return_indices=True)
alloc = a[tuple(idx)]
a0 = np.where(dist<=buff_dist, alloc, a) #0)
return a0
def shrink_(a, val=1, mask_vals=0, buff_dist=1):
"""Expand/buffer a raster by a distance
"""
if isinstance(val, (list, tuple)):
m = np.isin(a, val, invert=False).astype('int')
else:
m = np.where(a==val, 1, 0)
dist, idx = nd.distance_transform_edt(m, return_distances=True,
return_indices=True)
alloc = a[tuple(idx)]
m = np.logical_and(dist>0, dist<=buff_dist)
a0 = np.where(m, alloc, a) #0)
return a0
def regions_(a, cross=True):
"""currently testing regiongroup
np.unique will return values in ascending order
"""
if (a.ndim != 2) or (a.dtype.kind != 'i'):
msg = "\nA 2D array of integers is required, you provided\n{}"
print(msg.format(a))
return a
if cross:
struct = np.array([[0,1,0], [1,1,1], [0,1,0]])
else:
struct = np.array([[1,1,1], [1,1,1], [1,1,1]])
#
u = np.unique(a)
out = np.zeros_like(a, dtype=a.dtype)
details = []
is_first = True
for i in u:
z = np.where(a==i, 1, 0)
s, n = nd.label(z, structure=struct)
details.append([i, n])
m = np.logical_and(out==0, s!=0)
if is_first:
out = np.where(m, s, out)
is_first = False
n_ = n
else:
out = np.where(m, s+n_, out)
n_ += n
details = np.array(details)
details = np.c_[(details, np.cumsum(details[:,1]))]
return out, details
def aggreg_(a, win=(3,3), agg_type='mode'):
"""Aggregate an array using a specified window size and an aggregation
type
Parameters
----------
a : array
2D array to perform the aggregation
win : tuple/list
the shape of the window to construct the blocks
agg_type : string aggregation type
max, mean, median, min, mode, range, sum, central
"""
blocks = block(a, win=win)
out = []
for bl in blocks:
b_arr = []
for b in bl:
if agg_type == 'mode':
uni = np.unique(b).tolist()
cnts = [len(np.nonzero(b==u)[0]) for u in uni]
idx = cnts.index(max(cnts))
b_arr.append(uni[idx])
elif agg_type == 'max':
b_arr.append(np.nanmax(b))
elif agg_type == 'mean':
b_arr.append(np.nanmean(b))
elif agg_type == 'median':
b_arr.append(np.nanmedian(b))
elif agg_type == 'min':
b_arr.append(np.nanmin(b))
elif agg_type == 'range':
b_arr.append((np.nanmax(b) - np.nanmin(b)))
elif agg_type == 'sum':
b_arr.append(np.nansum(b))
elif agg_type == 'central':
b_arr.append(b.shape[0]//2, b.shape[1]//2)
else:
tweet("\naggregation type not found {}".format(agg_type))
b_arr.append(b.shape[0]//2, b.shape[1]//2)
out.append(b_arr)
out = np.array(out)
return out
You must be a registered user to add a comment. If you've already registered, sign in. Otherwise, register and sign in.